Get hands-on experience working with a data independent acquisition (DIA) parallel accumulation serial fragmentation (PASEF) data, from a Bruker timsTOF instrument, using a 3-organism mix data set created for instruction, based on the Navarro, Nature Biotech 2016 benchmarking paper. Use the Import Peptide Search wizard for DIA to build a spectral library from ddaPASEF data with automatic iRT calibration for retention time calibration and an mProphet learned model for peptide peak detection. Assess the data quality using a rich collection of Skyline summary plots including Retention Times, Peak Areas, Mass Accuracy, and CVs. Finally, perform a group comparison and make your own assessment of how well the data capture the expected ratios for each organism. (36 pages)
[pdf] [html] [small data] [full data]
* - written on v21.1, updated for v22.2
This is a companion to the Analysis of DIA/SWATH Data tutorial for Q Exactive and TripleTOF instruments.
[tutorial]
On December 7, 2021, the Skyline Team produced Webinar #21: Analysis of diaPASEF Data With Skyline, another great resource for this topic.
[webinar]
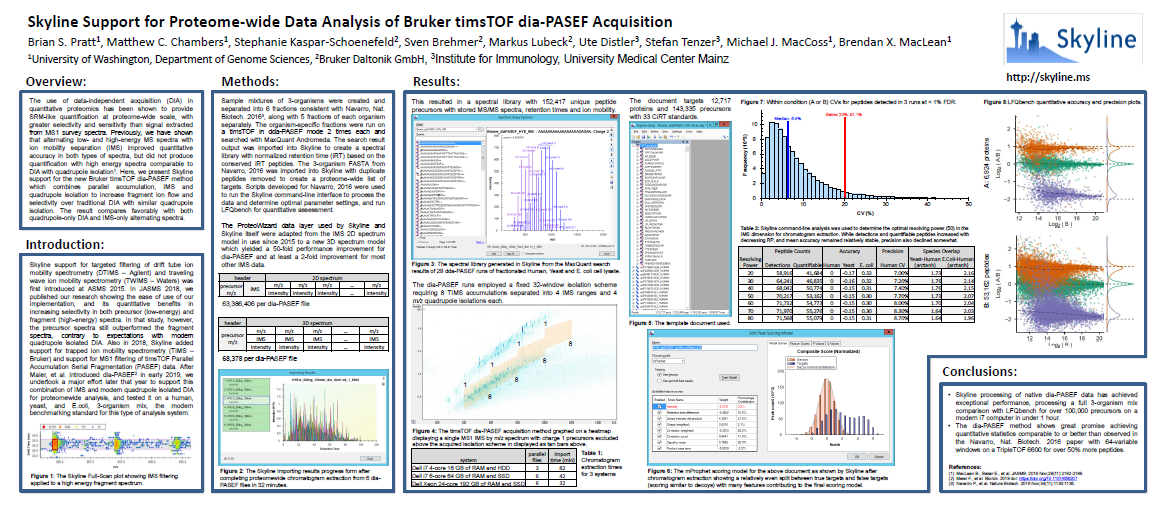
Learn more about about processing diaPASEF data with Skyline by reading the ASMS 2020 poster.
| previousnext |
| expand allcollapse all |